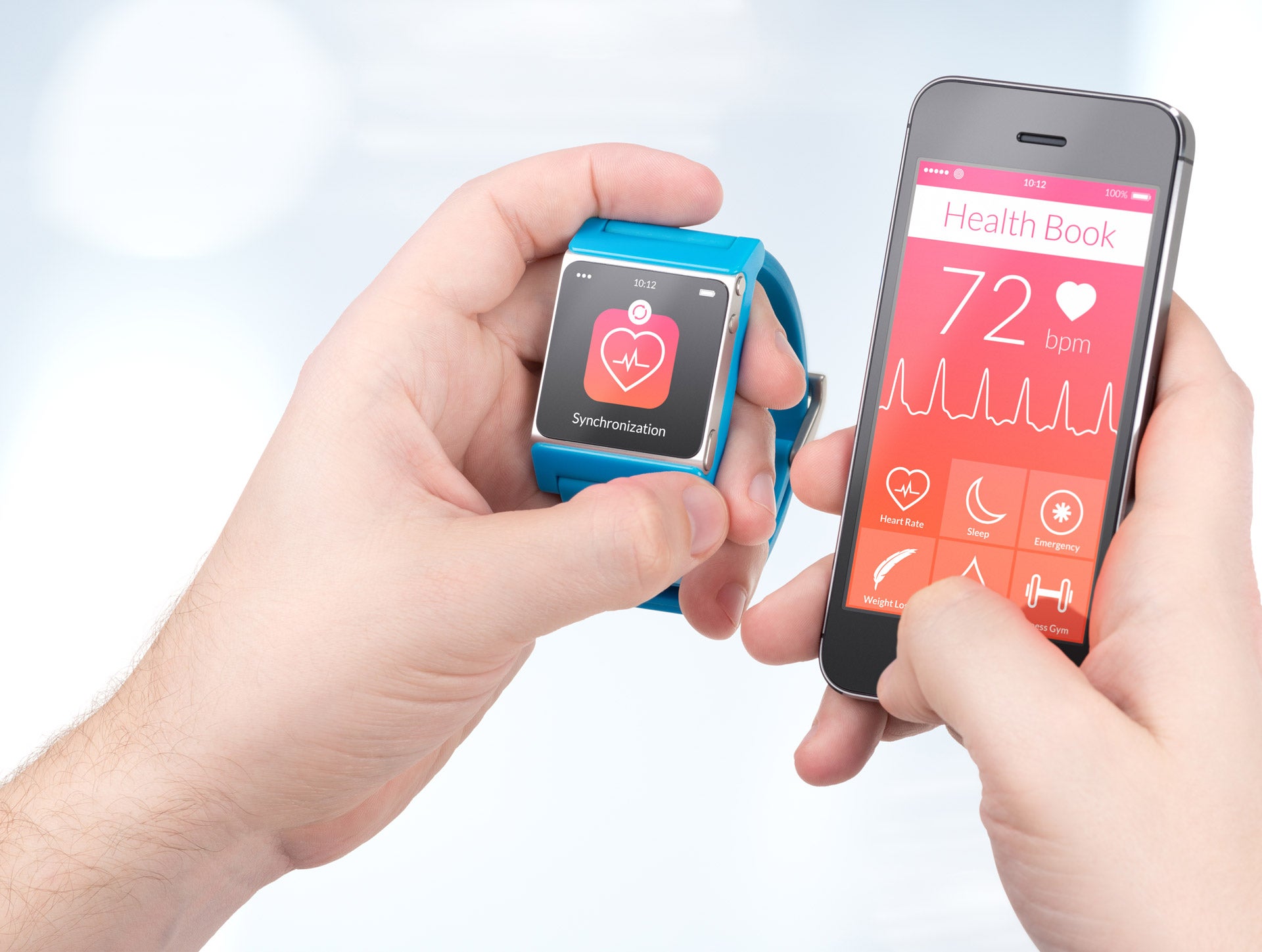
An eight-year study has revealed the benefits of continuous health monitoring – even when a person shows no signs of illness – suggesting a major paradigm shift is needed in healthcare.
Scientists at Stanford University School of Medicine and their collaborators studied 109 people using a combination of wearable technology, genome sequencing and microbial and molecular profiling.
Access deeper industry intelligence
Experience unmatched clarity with a single platform that combines unique data, AI, and human expertise.
This data gave a baseline health reading for the participants and allowed scientists to spot early signs of illness. A heart rate monitor, for example, was able to diagnose sleep apnea or indicate infection.
In all, the researchers detected 67 health alerts that could then be treated. These ranged from high blood pressure, arrhythmias, cardiomyopathy and early-stage cancer detection, among others. Many of the findings would have been missed in traditional health screenings.
“What this paper really shows is that if doctors and scientists do more advanced profiling reasonably frequently, they’ll discover clinically actionable information for patient health at a broader scale than has ever been shown before,” said Michael Snyder, Stanford University professor and chair of genetics.
Synder argues that current, more reactive, approaches in medicine are “deeply flawed” and could be “significantly improved” by with a continuous health monitoring approach.
Being proactive rather than reactive could prove a more cost-effective approach to healthcare – especially at a time when many health services, such as the UK’s NHS, are struggling with budget cuts and growing populations.
Building a biological picture with continuous health monitoring
Participants were tracked for an average of around three years, with some tracked for as long as eight years. Combining the data, which was also gathered from blood tests and stool samples, allowed the researchers to build a detailed understanding of participants’ biology.
“We caught a lot of health issues because we noticed their delta, or their change from baseline,” said Snyder. “For instance, we caught nine people with diabetes as it was developing by continuously monitoring their glucose and insulin levels.”
Genetic sequencing identified 13 disease-related findings. In one case, a young person had a mutation in a gene that put them more at risk for cardiomyopathy, which affects the heart’s muscles. Tests confirmed that the person did, in fact, have heart disease.
This data-led, continuous health monitoring approach is highly advantageous for patients, who can be alerted to a disease before showing symptoms and then receive treatment much earlier on.
However, genetic sequencing could have its downsides, such as creating information asymmetry in the health insurance industry. Should insurance providers have access to this data they could, in theory, use the data to increase premiums for people based on their likelihood to get a disease in later life.
Alternatively, if insurers did not have access to this data, patients could use it to take out insurance in the knowledge that they are more likely to require treatment for a particular disease.
More research needed to shift the paradigm
Snyder said the next step is increasing the number of participants being monitored.
The research could have also discovered new indicators, or biomarkers, for certain diseases. They will require further studies to confirm.
“Right now, we’re pretty much in the dark until we profile people, but this approach could provide us a much better view of people’s norms, what it means to be healthy and what it means when people deviate from that,” added Snyder.
“Ultimately, we want to shift the practice of medicine from treating people when they are ill to a focus on keeping them healthy by predicting disease risk and catching disease before it is symptomatic.”
The findings were published yesterday in the journal Nature Medicine.
Researchers from the Veteran Affairs Palo Alto Health Care System, St Vincent’s Hospital, University of Melbourne, University of California-Berkeley, Jackson Laboratory for Genomic Medicine, UC-San Francisco and UCLA contributed also to the research.
Read more: How virtual healthcare is targeting a “huge transformation” in Indian medicine







