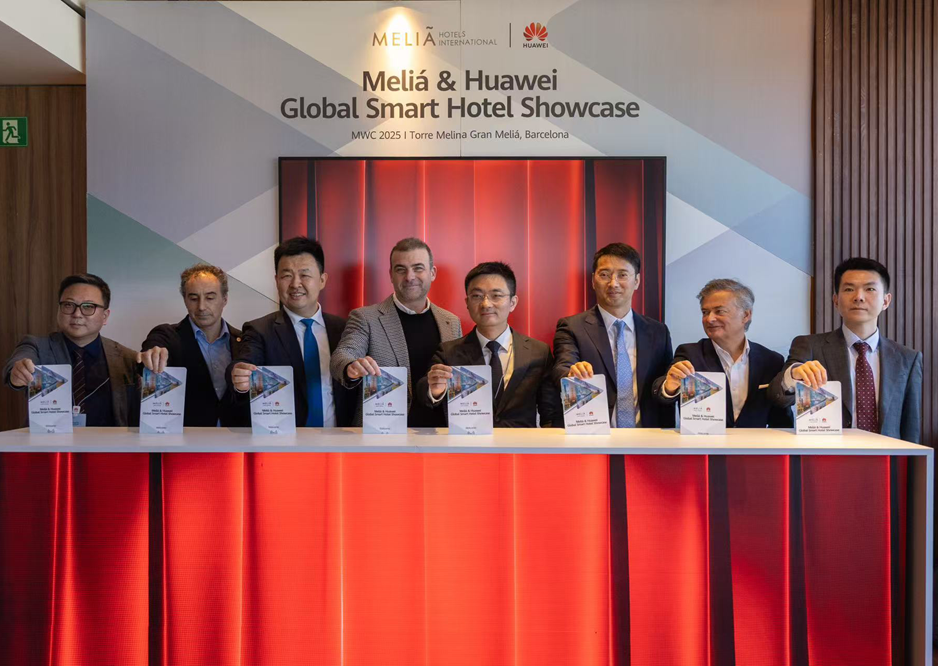
Researchers at the Francis Crick Institute have developed a database of gene activity in mice, which could reduce the use of animals in medical research.
According to the Foundation of Biomedical Research, 95% of animals used in scientific research are mice and rats, with 3.97 million experiments carried out on animals in the UK in 2017. Although the ethics of animal testing may be hotly debated, currently, when studying a particular disease, researchers create, infect, cull, obtain samples from mice and extract and sequence the RNA.
Access deeper industry intelligence
Experience unmatched clarity with a single platform that combines unique data, AI, and human expertise.
However the new database, which shows the immune response all of the genes from the blood of mice had to different pathogens, could prevent thousands of mice being used in individual experiments.
How the gene activity database could reduce mice testing
Accessible via an app, researchers will be able to view the activity of any gene across a range of diseases without needing their own mice. This could reduce the use of mice testing, saving money but also reducing the number of mice that have to be culled once experiments have finished.
Using sequencing technology, researchers measured gene activity across the different diseases and analysed the genetic signatures associated with these diseases to help understand the immune response. Crick group leader Anne O’Garra explains how this works:
“Gene activity can show us how the body responds to infections and allergens. There are thousands of genes involved in any immune response, so Akul Singhania, a Bioinformatics Postdoc, in our lab used advanced bioinformatics approaches to cluster the genes into modules.
“These modules represent clusters of genes that are co-regulated and can often be annotated to determine their function and known physiological roles. For example, of 38 lung modules there is a module associated with allergy, and seen only in the allergy model, containing over 100 genes and another module associated with T cells containing over 200 genes.”
“By sequencing both lung tissue and blood, we can also see how the immune response in the blood reflects the local response in the lung, and vice versa. This will help us to understand what we can learn from genetic signatures in the blood, since for most diseases doctors can’t realistically get lung samples from patients.”
Researchers anywhere in the world can now view this information on the gene activity in the lungs and blood of mice infected with a range of pathogens including the , influenza virus, the respiratory syncytial virus, and the house dust mite allergen.
This is the result of a ten-year project, with researchers first used a technique called microarray back in 2009 to detect gene activity in lung and blood samples.
O’Garra explains that since the project began, advances in this area, such as bioinformatics techniques, have meant that the database can be presented in a way that is accessible to anyone:
“Ten years since the project began, we now have an open access resource of gene expression that anyone in the world can use to look up their favourite genes and also see if they are regulated by type I or type II interferon signalling. Nobody said science was easy, but it’s certainly worthwhile.”
Read more: How close are 3D-printed organs to reality?







